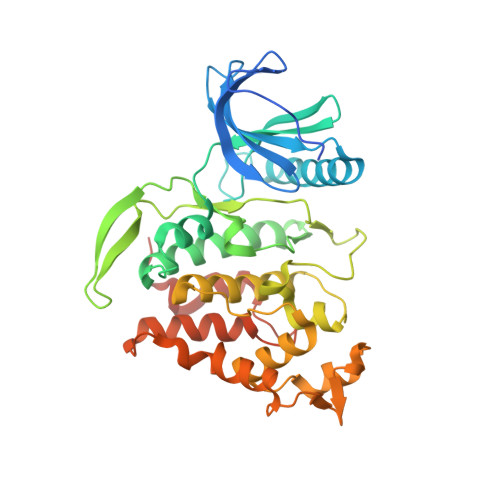
X-ray Structures and Feasibility Assessment of CLK2 Inhibitors for Phelan-McDermid Syndrome.
Kallen, J., Bergsdorf, C., Arnaud, B., Bernhard, M., Brichet, M., Cobos-Correa, A., Elhajouji, A., Freuler, F., Galimberti, I., Guibourdenche, C., Haenni, S., Holzinger, S., Hunziker, J., Izaac, A., Kaufmann, M., Leder, L., Martus, H.J., von Matt, P., Polyakov, V., Roethlisberger, P., Roma, G., Stiefl, N., Uteng, M., Lerchner, A.(2018) ChemMedChem 13: 1997-2007
- PubMed: 29985556
- DOI: https://doi.org/10.1002/cmdc.201800344
- Primary Citation of Related Structures:
6FYI, 6FYK, 6FYL, 6FYO, 6FYP, 6FYR, 6FYV - PubMed Abstract:
CLK2 inhibition has been proposed as a potential mechanism to improve autism and neuronal functions in Phelan-McDermid syndrome (PMDS). Herein, the discovery of a very potent indazole CLK inhibitor series and the CLK2 X-ray structure of the most potent analogue are reported. This new indazole series was identified through a biochemical CLK2 Caliper assay screen with 30k compounds selected by an in silico approach. Novel high-resolution X-ray structures of all CLKs, including the first CLK4 X-ray structure, bound to known CLK2 inhibitor tool compounds (e.g., TG003, CX-4945), are also shown and yield insight into inhibitor selectivity in the CLK family. The efficacy of the new CLK2 inhibitors from the indazole series was demonstrated in the mouse brain slice assay, and potential safety concerns were investigated. Genotoxicity findings in the human lymphocyte micronucleus test (MNT) assay are shown by using two structurally different CLK inhibitors to reveal a major concern for pan-CLK inhibition in PMDS.
Organizational Affiliation:
Novartis Institutes for BioMedical Research, Novartis Campus, 4002, Basel, Switzerland.